Finding the proper chemical compound for a specific application, while maintaining development costs to the lowest possible level, can be difficult. Indeed, to find a suitable candidate, modern drug discovery approaches often comprise screening a staggering amount of chemicals from a library and conducting iterative cycles of compound optimization. Subsequently, success in clinical trials is not guaranteed, with ~40% of drugs in phase III trials failing at this stage (1) and causing serious financial and time losses for the companies and stakeholders investing into new chemical entities development. It is therefore important to invest sufficiently and plan carefully the preclinical stages of development, to rapidly rule out harmful or ineffective compounds and identify the best candidates. Typically, preclinical studies consist of an initial in vitro screening of biological activity and toxicity in cell lines, followed by in vivo assays, usually executed in rodents. This transition from cell-based assays to a whole complex organism often does not proceed without issues, and multiple discrepancies between the two models have been documented (2, 3). Such poor predictivity at the stage of in vitro assays could result in the selection of ineffective or toxic compounds for the subsequent expensive trials on mice or the elimination of good leads from the pipeline due to a toxic effect observed exclusively in vitro, but non-existent in vivo! These fruitless trials are also inflicting unneeded suffering upon the test animals – while in the 3Rs (Replace, Reduce, Refine) era (4) each compound should be profiled to the best extent possible before testing it on mammals.
As we have seen, an accurate characterization of the candidate compounds at the early research stages is crucial to avoid time- and money-costly failures at later stages. A solution to bridge the gap between cell-based assays and tests performed in rodents and humans would be models more complex than cells or tissues, but easier to manipulate than rodents. C. elegans perfectly fits such requirements: it is a valuable tool in toxicology and can predict toxicity outcomes in mammals (5, 6). Additionally, the use of C. elegans is devoid of ethical concerns and its short lifecycle and lifespan guarantee that results can be obtained rapidly.
Figure – Proposed outline of a compound testing pipeline
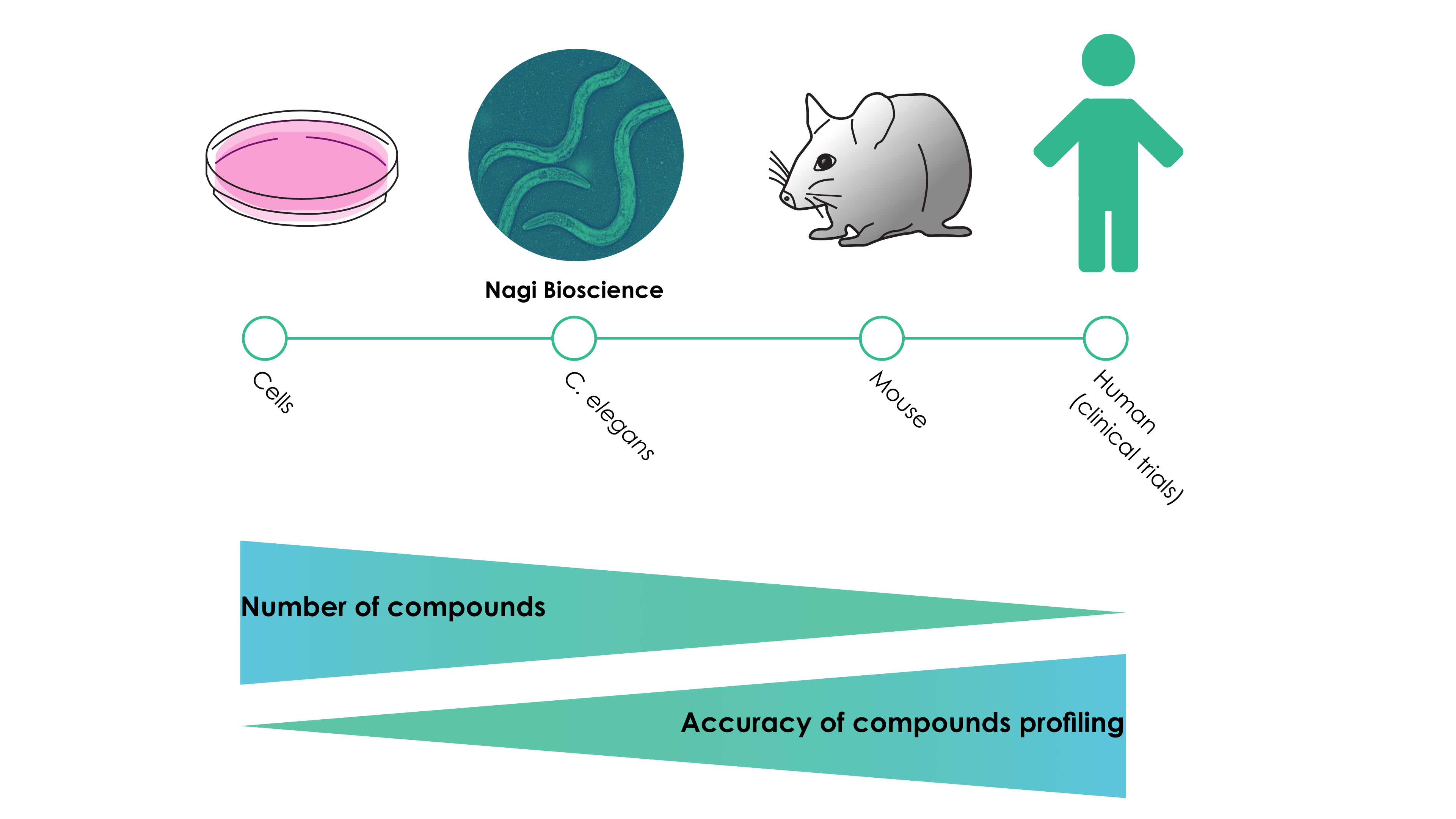
Until now, however, the use of C. elegans for large-scale studies has been limited by the fact that the maintenance of the worms and the data acquisition required extensive manual handling. This implies that for large-scale projects trained specialists would have to be employed en masse. At Nagi Bioscience, we bypassed this roadblock by creating an automated high-throughput testing platform that requires minimal handling by the operator. Our platform hence significantly decreases the amount of man-hours spent on obtaining relevant biological data. Importantly, the developed platform can facilitate different types of assays (see the applications page on our website), offering an ideal assay for various types of projects.
References
1. Clinical Development Success Rates 2006-2015—BIO, Biomedtracker, Amplion 2016.pdf. (n.d.). Retrieved 21 September 2021, from https://www.bio.org/sites/default/files/legacy/bioorg/docs/Clinical%20Development%20Success%20Rates%202006-2015
%20-%20BIO,%20Biomedtracker,%20Amplion%202016.pdf
2. Saeidnia, S., Manayi, A., & Abdollahi, M. (2015). From in vitro Experiments to in vivo and Clinical Studies; Pros and Cons. Current Drug Discovery Technologies, 12(4), 218–224. https://doi.org/10.2174/1570163813666160114093140
3. Bowman, C. M., & Benet, L. Z. (2019). In Vitro–In Vivo Inaccuracy: The CYP3A4 Anomaly. Drug Metabolism and Disposition, 47(12), 1368–1371. https://doi.org/10.1124/dmd.119.088427
4. https://nc3rs.org.uk/the-3rs
5. Leung, M. C. K., Williams, P. L., Benedetto, A., Au, C., Helmcke, K. J., Aschner, M., & Meyer, J. N. (2008). Caenorhabditis elegans: An Emerging Model in Biomedical and Environmental Toxicology. Toxicological Sciences, 106(1), 5–28. https://doi.org/10.1093/toxsci/kfn121
6. Williams, P. L., & Dusenbery, D. B. (1988). Using the nematode Caenorhabditis elegans to predict mammalian acute lethality to metallic salts. Toxicology and Industrial Health, 4(4), 469–478. https://doi.org/10.1177/074823378800400406